Tags:
Define-XMLOne of the most widely used standards today is Define-XML. The latest version of Define-XML is v2.1, which went live in May 2019. Get release information on the CDISC website.
What is Define-XML?
According to CDISC: “Define-XML is required by the United States Food and Drug Administration (FDA) and the Japanese Pharmaceuticals and Medical Devices Agency (PMDA) for every study in each electronic submission to inform the regulators which datasets, variables, controlled terms, and other specified metadata were used.”
The FDA’s Technical Conformance Guide explains that Define-XML is "arguably the most important part of the electronic dataset submission for regulatory review” because it helps reviewers gain familiarity with study data, its origins and derivations.
The standard itself is known as ‘Define-XML’. The file that’s submitted to the FDA upon completion is the data definition file, known simply as ‘define.xml’.
Define-XML as a dataset descriptor
It is commonly thought that Define-XML is simply a dataset descriptor: a way to document what datasets look like, including the names and labels of datasets and variables, what terminology is used etc. This is essentially what Define-XML was created for.
But by instead thinking of Define-XML as a tool to create better quality, more efficient clinical studies, users can unlock the true potential of the standard.
Progressive uses of Define-XML
You can use Define-XML to help you optimize the end-to-end clinical trial process in the following ways:
1) Use Define-XML in your CRF design process
Many organizations treat Define-XML as an afterthought: only when case report forms (CRFs) are designed, data is collected and the study is complete do they think about creating the define.xml file for FDA submission.
But this approach can lead to incomplete data, the need for protocol amendments, complex mapping, increased quality control. How do you know when designing the CRF that you’re collecting all the relevant SDTM data? And when the data has been collected, how can you verify the submission is what you intended when you have no study definition to compare it with? It can take valuable time and resources to make sure all data has been collected in the right format, and ultimately can elongate the study process.
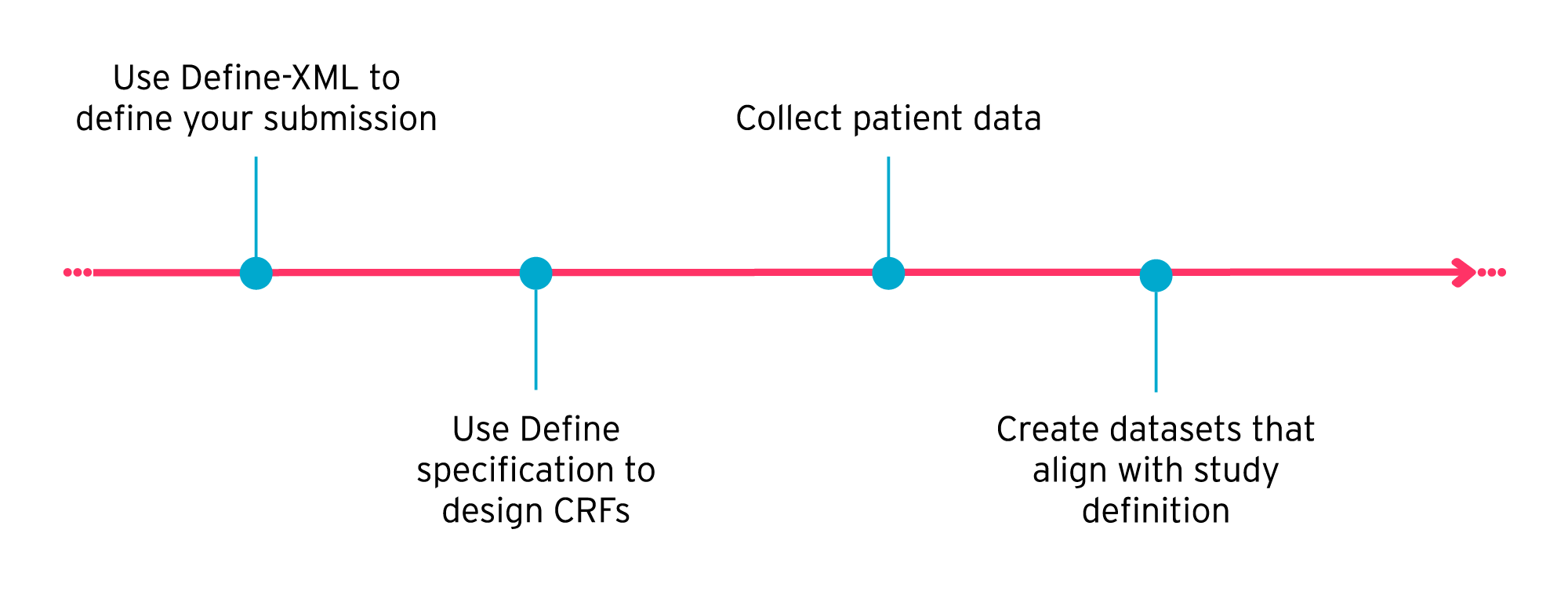
A more efficient approach is to use Define-XML to define your study, end-to-end, right at the start. This includes defining SDTM, SEND, and ADaM datasets upfront.
 Using Define-XML and SDTM to design submission datasets at the start of a study makes it easier to set up your study and create your case report forms (CRFs). By setting out what information should ultimately appear in your submission datasets before you collect any patient data, you can create CRFs with confidence, knowing that you’re collecting all the required information in the right format.
Using Define-XML and SDTM to design submission datasets at the start of a study makes it easier to set up your study and create your case report forms (CRFs). By setting out what information should ultimately appear in your submission datasets before you collect any patient data, you can create CRFs with confidence, knowing that you’re collecting all the required information in the right format.
For example, the SDTM standard gives the ‘Identifier’, ‘Topic’, ‘Qualifier’, and ‘Timing’ variables required in your submission datasets. If you know upfront what variables to use, you can create your CRFs accordingly.
You can also do your dataset annotation of CRFs with SDTM variables upfront. This can help ensure all your collected data has a place in SDTM. This has the additional benefit of providing basic mapping between the forms and the datasets. CDISC provides a mechanism to extend Define-XML which is permissible and allows the storage of additional metadata such as complex dataset mappings (e.g. how data may be merged into one single dataset from two sources).
In this way, using Define-XML upfront, rather than retrospectively, can help you ensure your study is a success. Read more about using Define-XML for dataset design.
2) Use Define-XML in EDC data conversions
Define-XML is not limited to just describing CDISC SDTM and ADaM dataset structures. From an electronic data capture (EDC) system, you can export proprietary dataset formats which can be described using the Define-XML model. With the right tools, you can automatically generate a Define-XML that describes the EDC export datasets using the CRFs/eCRFs themselves. This can then be displayed in a friendly HTML or PDF format allowing early visibility of the datasets that will be delivered by the EDC system.
The Source Proprietary dataset spec enables upfront mapping of EDC datasets to SDTM datasets. These mappings can be described (and made machine-executable) using extensions to Define-XML and human-readable SDTM mapping specifications produced automatically, aiding review and approval of mappings.
In addition, the Define-XML mapping extensions provide a machine-executable format that can be processed by data transformation code to enable the automatic conversion of datasets in commercially available tools.
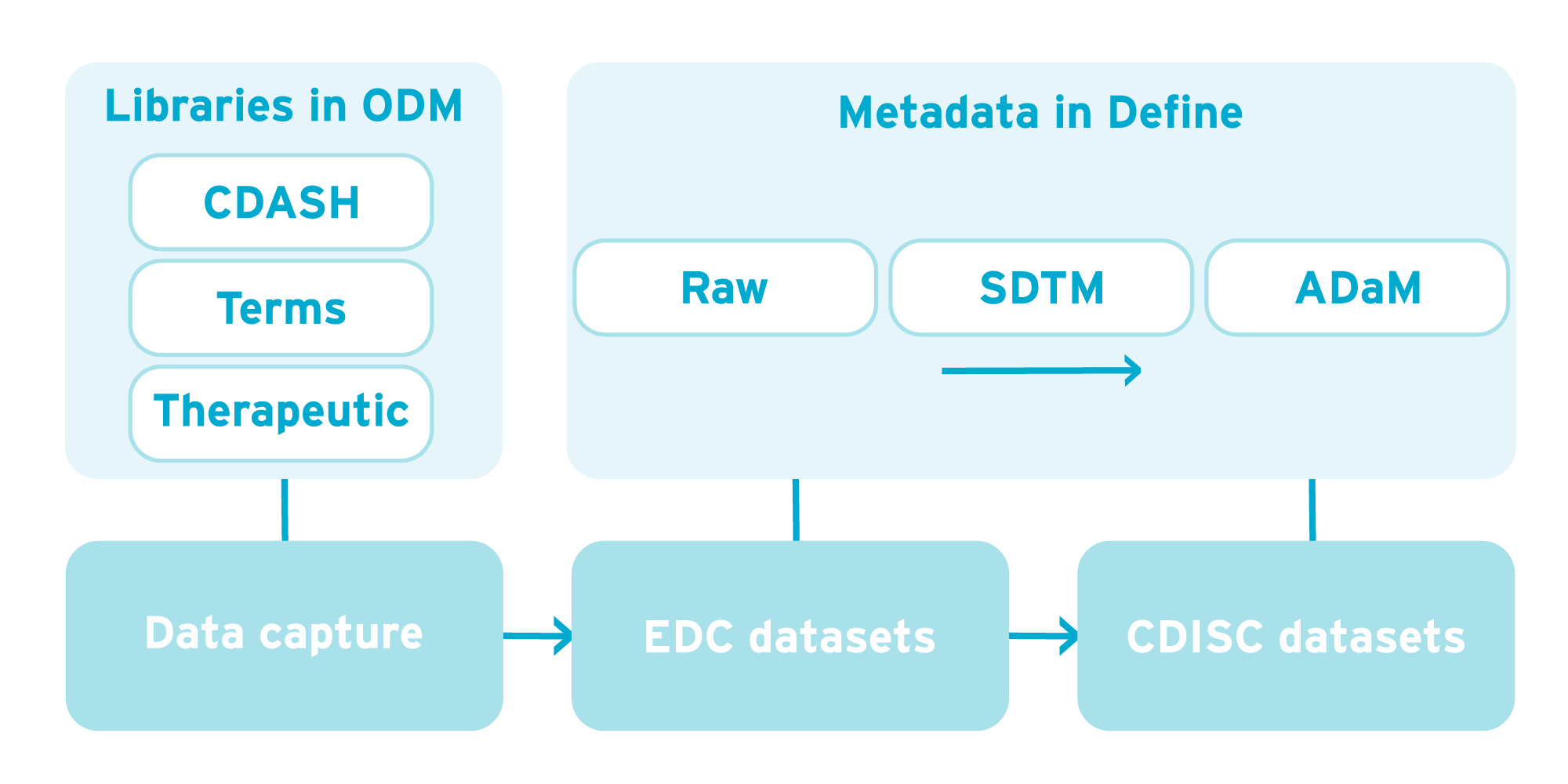
The diagram below shows the flow of data from data capture through to CDISC datasets and the part CDISC metadata plays. Metadata is used in designing data capture forms using CDISC ODM and Define-XML in designing destination datasets. All of this vendor-neutral metadata can form the basis of form and dataset libraries which can be re-used from study to study.
3) Creating and re-using dataset libraries
Define-XML is the perfect tool to help you create libraries of datasets (EDC, SDTM, ADaM), mappings, page links to CRF variables, and so on for re-use from one study to the next.
A metadata-driven approach using Define-XML can optimize a single study from set-up to submission. But creating libraries of reusable metadata will make future studies even more efficient.
If you have a library of data acquisition forms, proprietary EDC datasets, SDTM datasets, ADaM, and dataset mappings that are approved internally and ready to use, you’ll only have to create new content where there is a specific requirement for it. All other approved metadata is already there in your library.
4) Automating dataset validation
Another major advantage to defining datasets upfront is that validation can also be done up front. By creating a prospective definition of the intended datasets at the start of the study, it is possible to machine-validate study dataset designs for conformance to external standards. It is also possible to validate that populated datasets match the original specifications. This way, data quality and submission compliance are built-in upfront with less reliance on downstream validation.
We go into a little more detail on validation possibilities below:
- Compare study dataset designs, including controlled terminology, to external and internal standards
When designing SDTM datasets and creating controlled terms, it is imperative that these comply with the latest and/or chosen version of National Cancer Institute Controlled Terminology (NCI CT). During the dataset design phase, automatic comparisons and compliance checks should be made with the appropriate version of NCI CT.
Companies should also develop their own domains that comply with CDISC SDTM but include content that falls outside of the standard Implementation Guide domains. For example, specialist findings domains may be required for a particular therapeutic area. In this scenario, companies should compare study dataset designs against their own data standards to check for differences and either accept or reject them accordingly. - Compare ‘As specified’ study dataset specification against ‘As delivered’ study dataset designs
Increasingly, studies are outsourced to Contract Research Organizations (CRO) and this leads to an increased burden on sponsors. This tends to happen in two areas: (a) upfront specification of deliverables and (b) downstream validation of those deliverables.
When dataset validation is done upfront, a human-readable target SDTM specification (in HTML, PDF, Word or Excel) can be given to a CRO to describe what is expected to be in the delivered datasets: an ‘As specified’ study dataset specification.
When CROs return the datasets, they should also provide ‘As delivered’ study dataset metadata. With both ’As specified’ and ’As delivered’ study dataset metadata available, it is easy to compare the study dataset metadata to verify that the ’As delivered’ dataset actually matches what was specified. - Compare dataset data to dataset metadata and SDTM or ADaM
Having a target SDTM Define-XML available upfront allows automated comparison of delivered datasets against study dataset metadata, either as specified or as delivered. Comparing data to as specified Define-XML verifies that the data matches what was originally intended/specified. And comparing data to as delivered Define-XML ensures that the data matches the dataset definition. This is important as it will ultimately be this as delivered Define-XML that is submitted to the FDA.
You can also compare with CDISC standards such as SDTM and ADaM using the CDISC Open Rules Engine (CORE). CORE is a validation engine that can help you easily validate against the standardized conformance rules published by CDISC. In April 2023, we released Formedix CORE, a free desktop application for validating datasets using the CORE engine. This will validate against both SDTM/ADaM, and the Define-XML metadata. CDISC will also be adding the ability to validate against regulatory rules (FDA, PMDA etc) in the future.

Define.xml file submission
As we’ve shown, there are many benefits to using Define-XML - not only as a dataset descriptor, but as a means to streamline the clinical study process.
Define-XML should not be thought of as simply a submission deliverable, but as a CDISC model that helps optimize the end-to-end clinical trial process. It can be used to establish dataset libraries that promote study-to-study re-use, as well as to drive efficiencies through expedited study set-up and streamlined dataset conversions.
Learn how you can create Define XML in one click with our Visual Define.XML editor or check out of blog to what's new in Define-XML 2.0 including define xml 2.0 examples.
.
The 6 dos and don’ts of Define-XML
Ready to find out more about Define-XML creation?
Download our free guide to the 6 dos and don’ts of Define-XML to help you implement the standard and get your study submission ready.
Author's note: this blog post was originally published in May 2014 and has been updated for accuracy and comprehensiveness.
![]()
About the author

Kevin Burges
Head of Product Management | Formedix
Kevin Burges has been working at Formedix for over 20 years. Over time his role has changed from Developer to Senior Developer, to Technical Director and now Head of Product Management.
Kevin has a strong interest in metadata management and automation as an engine for streamlining clinical trials, and he works closely with customers to evolve the ryze platform with their needs in mind. He has also worked closely with CDISC since 2000, and has won awards for outstanding achievement towards advancing CDISC standards.
Nowadays, he’s part of the Data Exchange Standards team, which includes ODM, Define XML and Dataset XML.









